Tutorials
Before you start
Pre-analyzed example data can be found in the tutorials folder of the GitHub repository.
Before we analyze your own data, you can try to reproduce the results of these examples to get familiar with the workflow of OpenMebius2.
Installation
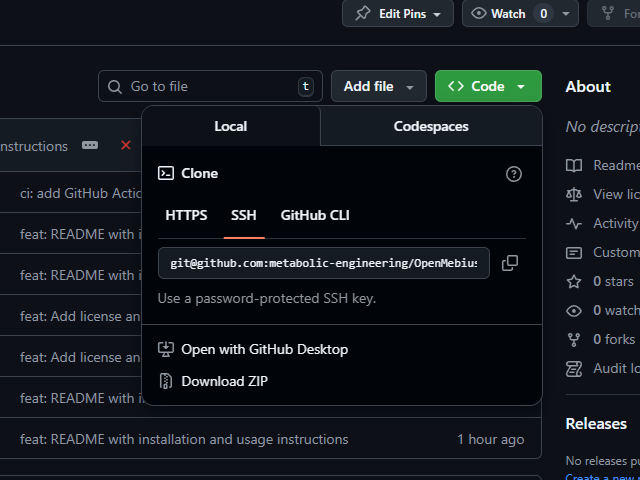
- Go to the GitHub repository and download the repository as a ZIP file by clicking the green
Codebutton and selectingDownload ZIP. If you are familiar with Git, you can also clone the repository by using the following command:git clone https://github.com/metabolic-engineering/OpenMebius2.git
- Unzip the downloaded ZIP file and navigate to the
OpenMebius2/installerfolder. This folder contains the installer for OpenMebius2. Click the installer file (depending on your operating system) to install OpenMebius2 on your computer (administrative privileges may be required). - Start OpenMebius2 program by double-clicking the OpenMebius2 icon on your desktop or in your applications folder.
Opening example projects
- Start OpenMebius2 program.
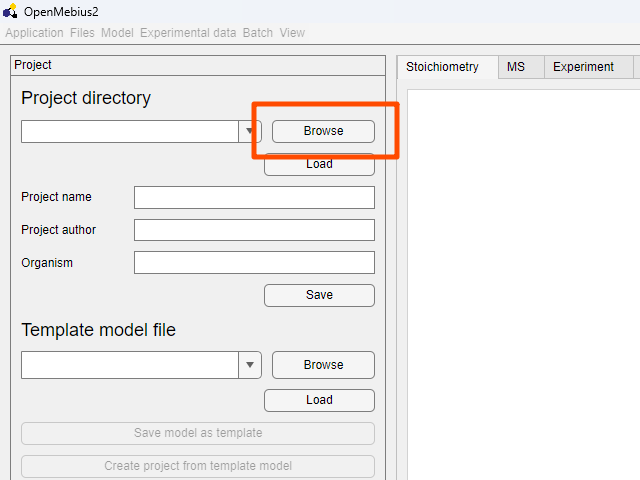
- Click the
Browsebutton in theProjectfield on the top left of the window.
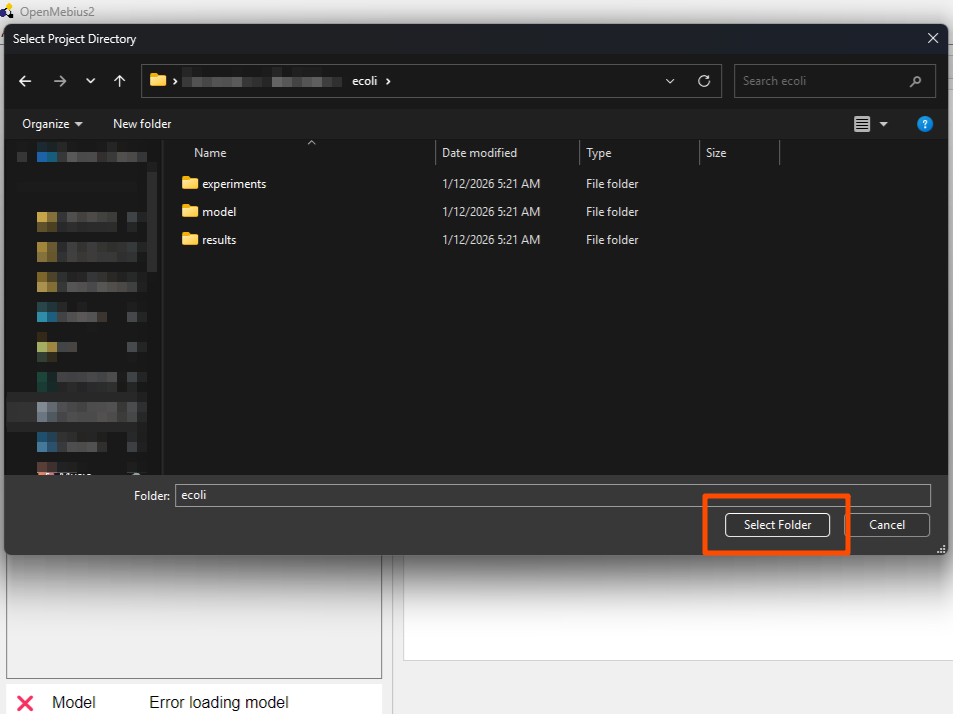
- Navigate to the
tutorials/ecolifolder in the unzippedOpenMebius2directory. Click theSelect Folderbutton to open the example project.
- Click the
Loadbutton to load the example project.
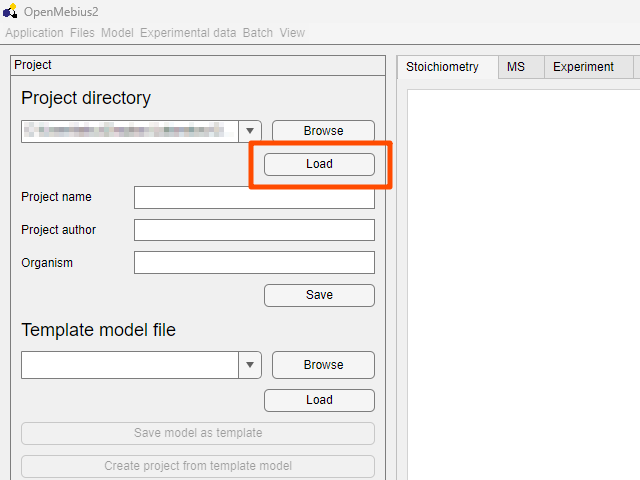
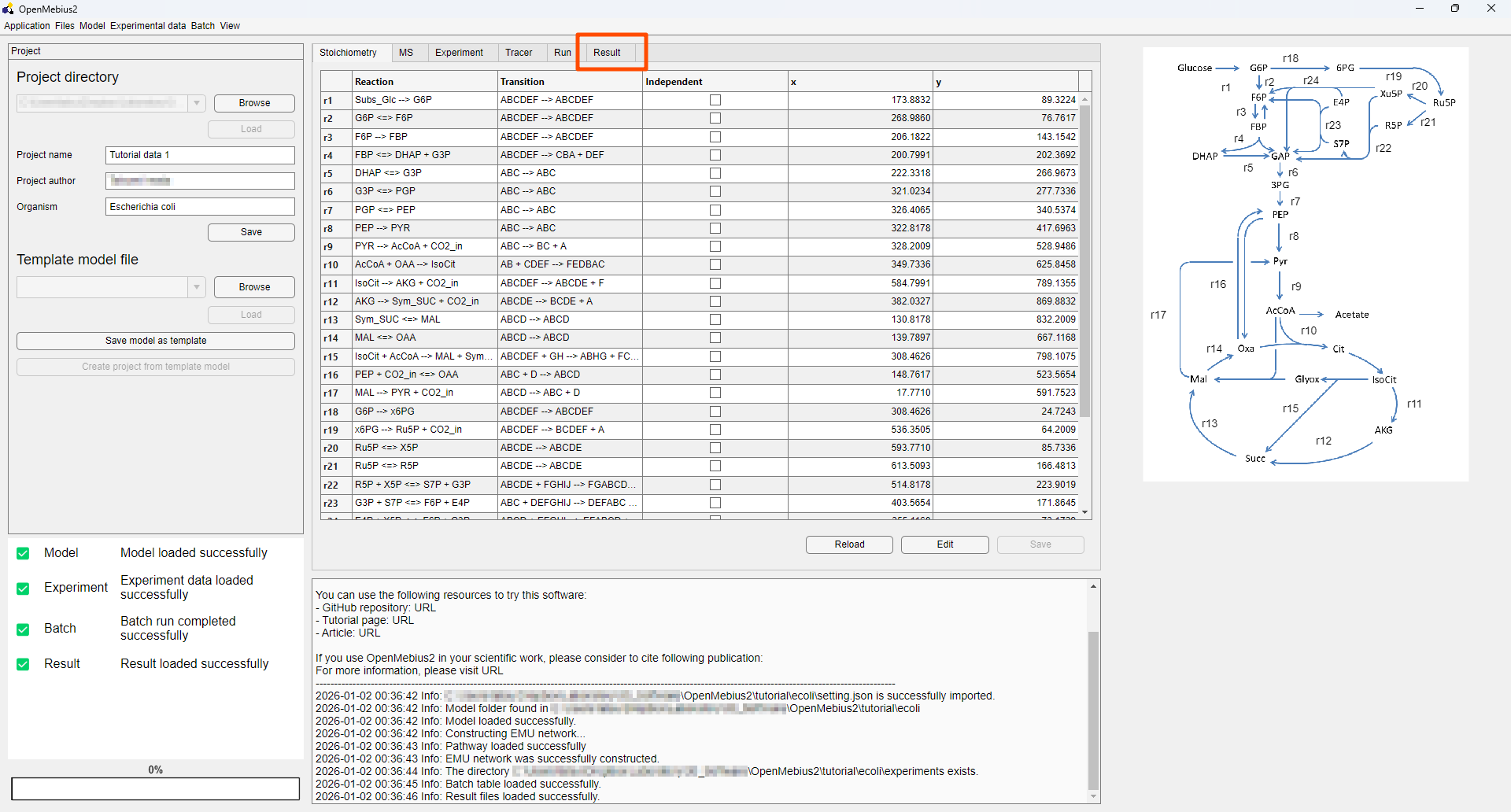
- The example project is now loaded, and you can select
Resulttab to view the analysis results.
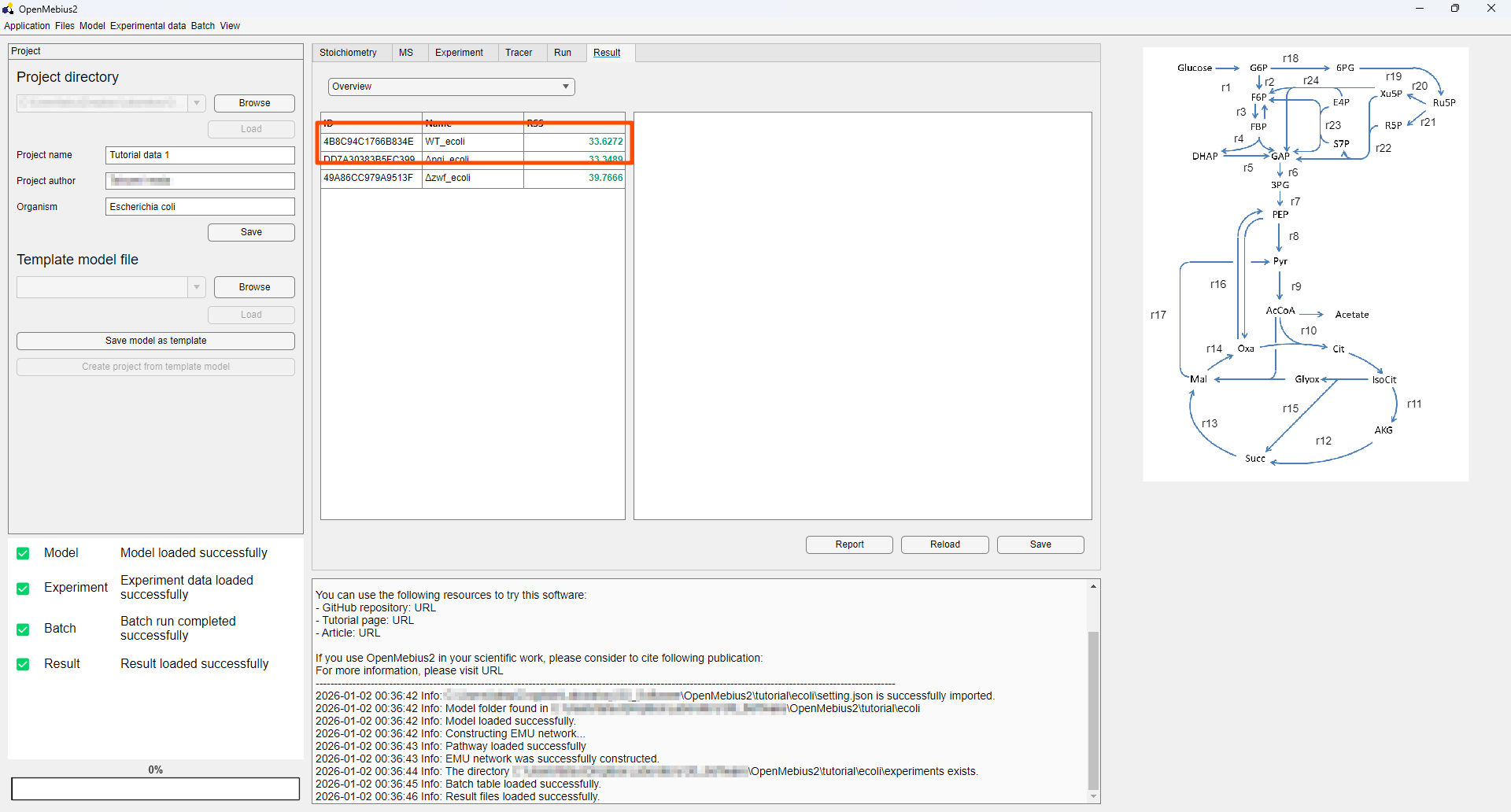
- By clicking the desired row in the result table, you can view fluxes and upper/lower bounds calculated by flux variability analysis (FVA).
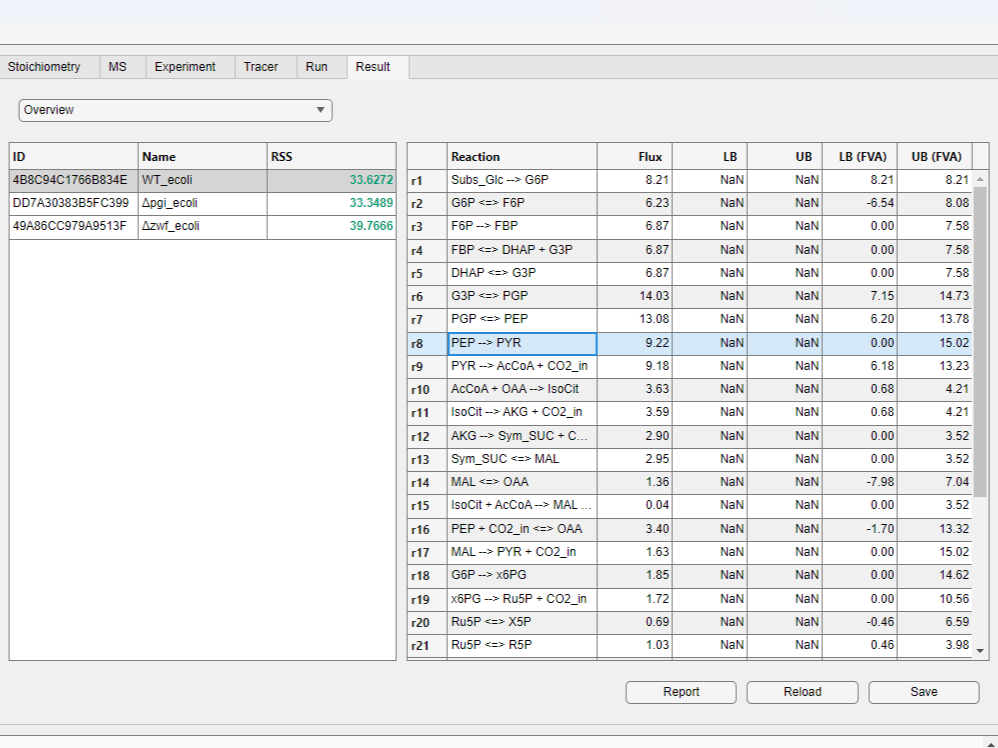
- Choose the
Detailsitem in the drop-down menu on the top of the result table to view simulated and measured mass distribution vectors (MDVs) for each metabolite.
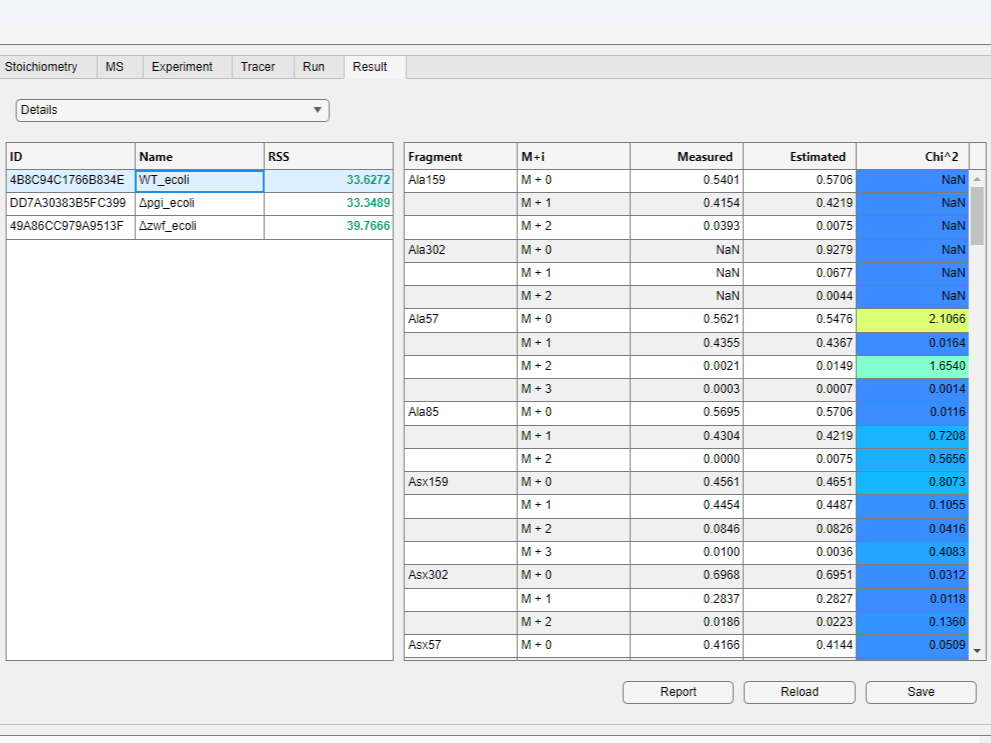
- You can compare fluxes of different results by selecting multiple rows in the result table while holding the
Ctrlkey (orCmdkey on Mac) and clicking theComparisonbutton above the table.
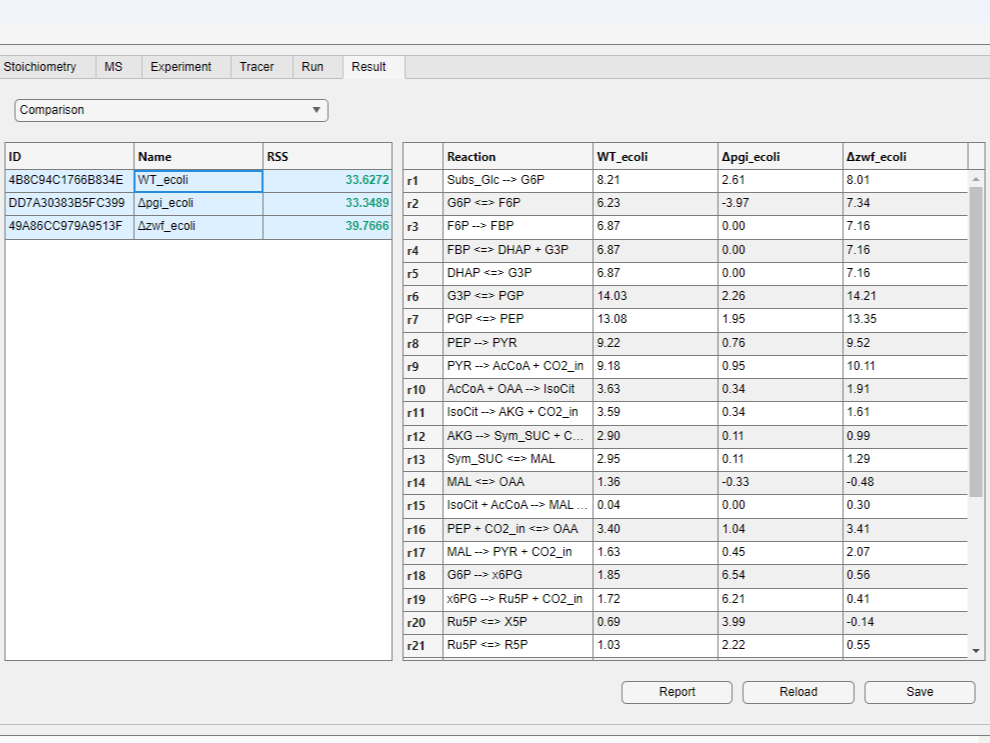
- You can also visualize flux maps on the right side of the window.
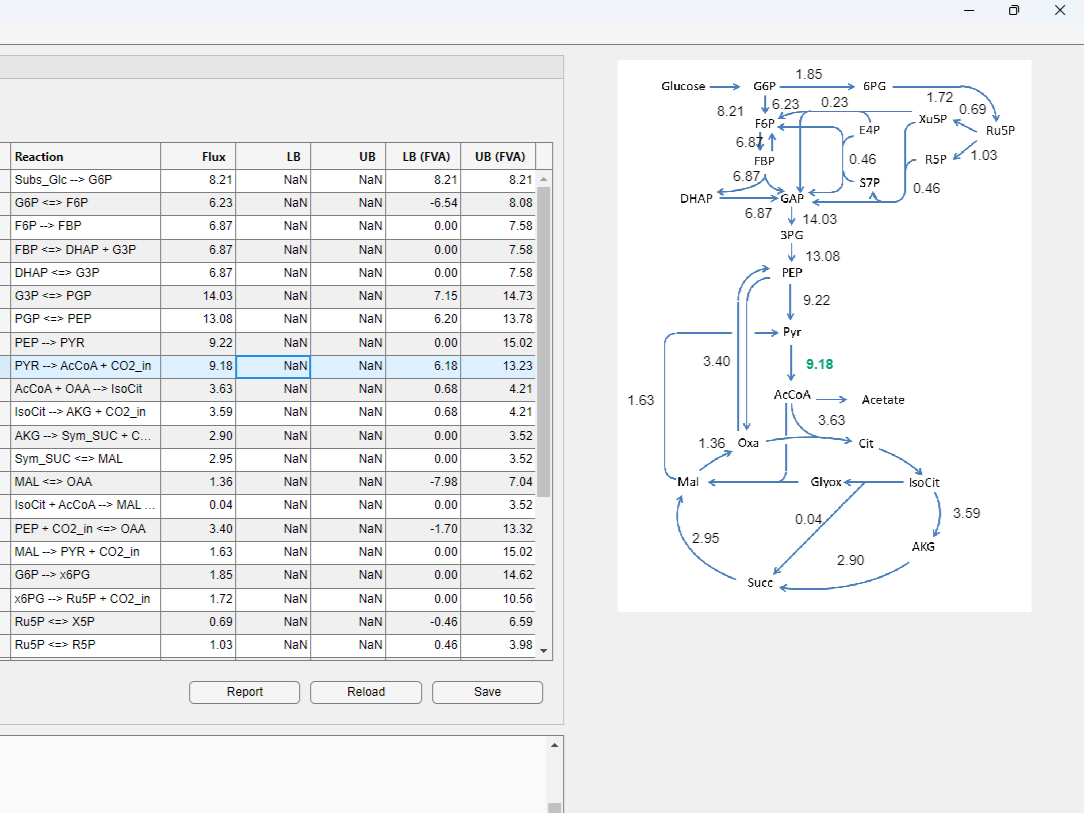
Create a new project
- Start OpenMebius2 program.
- Click the
Browsebutton in theTemplate Modelfield on the bottom left of the window.
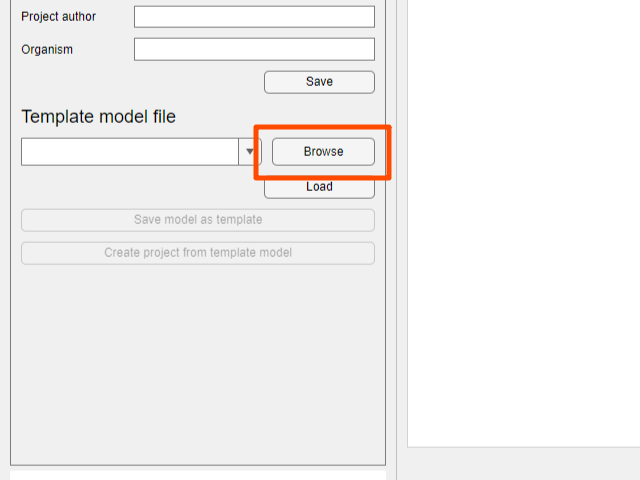
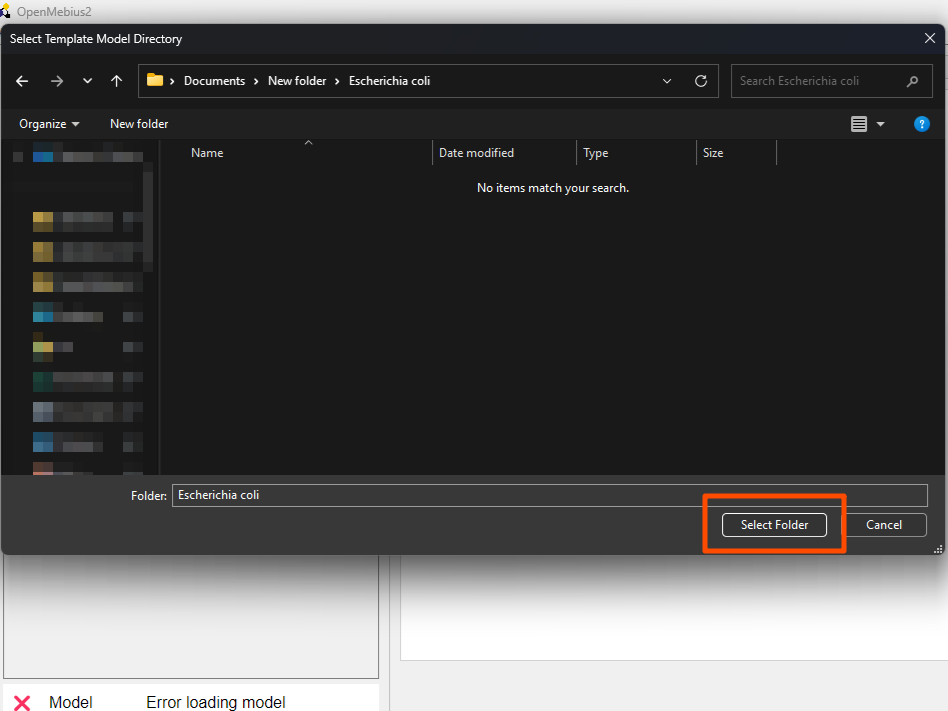
- Navigate to the
modelfolder in the unzippedOpenMebius2directory. Click theSelect Folderbutton to select the template model.
- Click the
Loadbutton to load the template model.
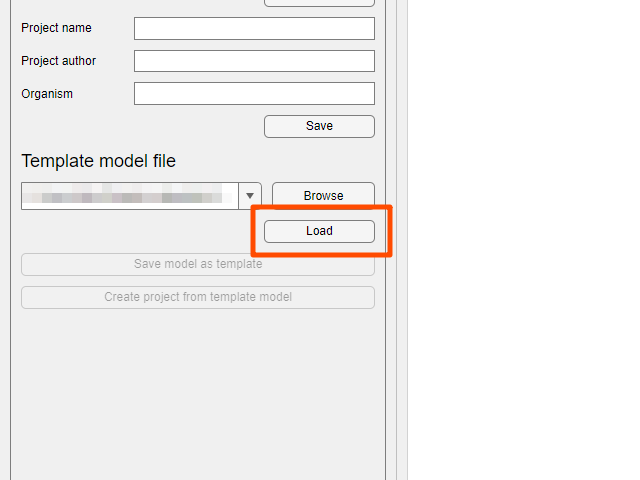
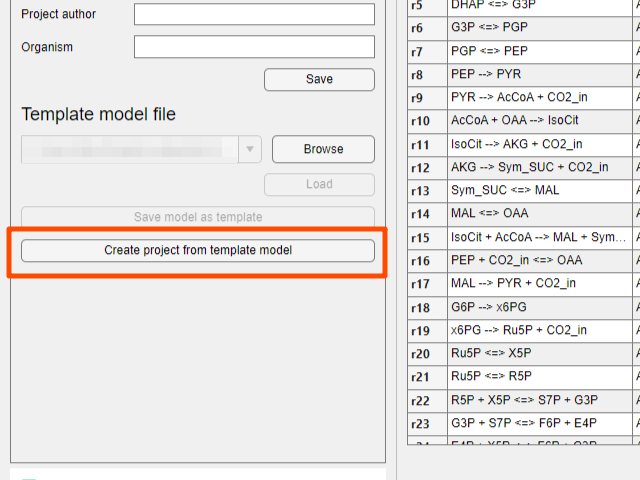
- After loading the template model, click the
Create project from template modelbutton to create a new project.
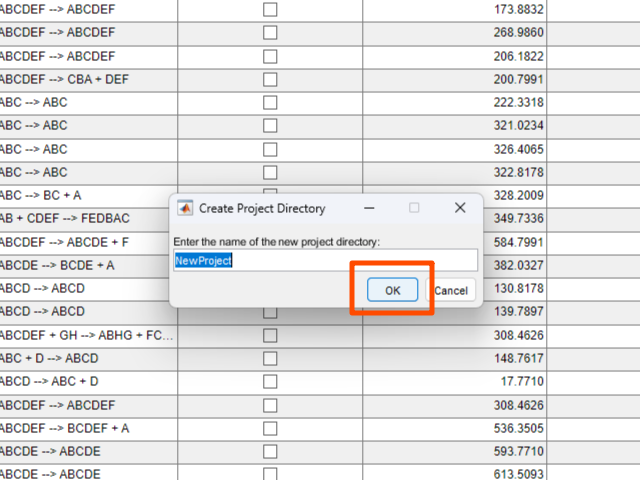
- In the dialog that appears, enter the project name. Then click the
OKbutton.
- Select the folder where you want to save the new project and click the
Select Folderbutton.
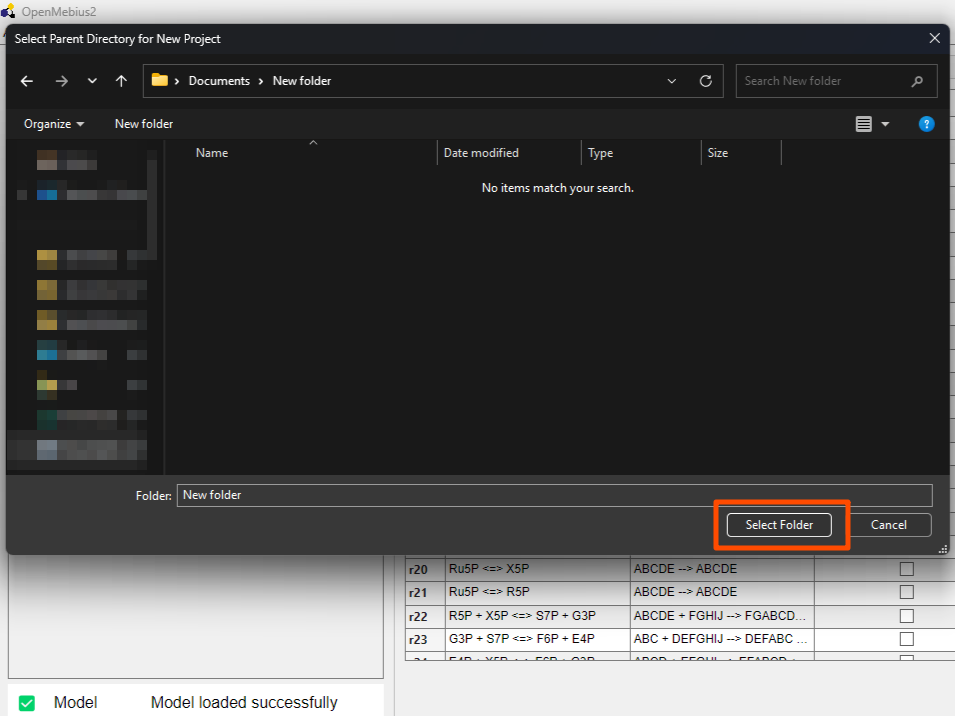
- Wait a moment while the new project is being created. The indicator icon on the model will turn green when the process is complete.
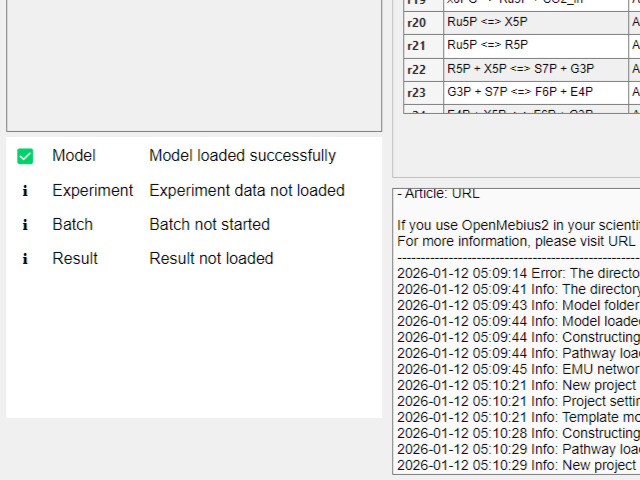
- The new project is now created and loaded. You can start customizing the model, adding experimental data, and performing analysis.