Data format
Model data format
Metabolic model consists of an Excel file defining the metabolic network, metabolic_model.xlsx, a label.json file defining labeling information, and an image file of the metabolic map, metabolic_pathway.tiff.
Each file is described below.
Metabolic model Excel file
The metabolic model Excel file consists of these sheets, info, model, MS, atom, biomass, and position.
| Sheet name | Description |
|---|---|
info |
General information about the model |
model |
Definition of reactions in the metabolic model |
MS |
Observed mass fragments for metabolites |
atom |
The number of atoms for each element in the metabolites |
biomass |
The biomass coefficients for the biomass equation |
position |
Positions of reactions in the metabolic map image |
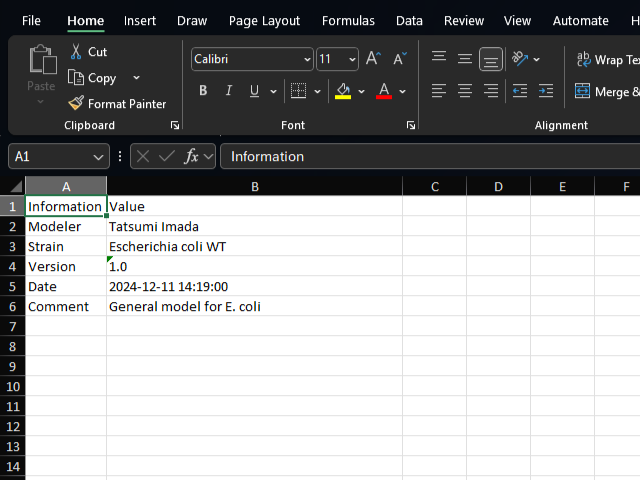
info sheet
The info sheet contains general information about the model, such as the model name, author, date, and description.
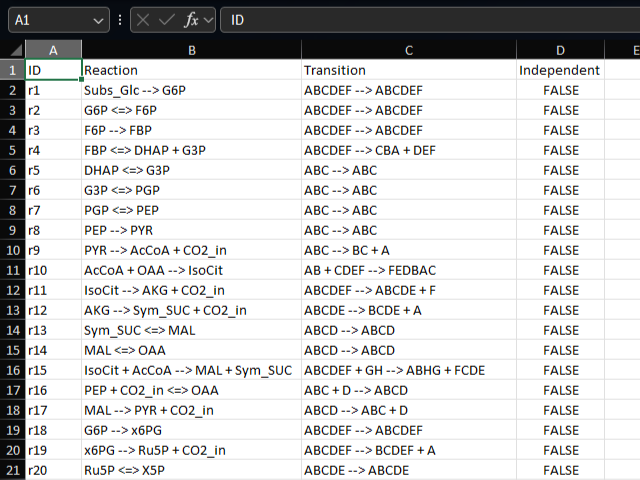
model sheet
The model sheet defines the reactions in the metabolic model. Each row represents a reaction, with columns for reaction ID, reaction, and atom mapping information.
Each reaction is represented in the format “A + B –> C + D”, where A, B, C, and D are metabolite IDs.
Reaction format
| Symbol | Meaning |
|---|---|
| (space) | Separates each metabolite and symbol. |
+ |
Indicates multiple metabolites. |
--> |
Indicates the irreversible reaction. |
<=> |
Indicates the reversible reaction. |
| Metabolite name | Meaning |
|---|---|
A |
Indicates metabolite A. |
Subs_A |
Indicates substrate A and this metabolite will become efflux reaction in the model. |
Sym_A |
Indicates symbiotic metabolite A such as a succinate. |
| Column name | Description |
|---|---|
ID |
Reaction ID, that must be unique in the model. |
Reaction |
Reaction formula as described above. |
Transition |
Atom mapping information for the reaction. The number of parts must match that of the reaction. Each part describes the mapping from substrate to product. |
Independent |
Boolean value indicating whether the reaction sets as independent (TRUE) or dependent (FALSE) in the model. If TRUE, the reaction flux will be enforced to be independent variable. |
MS sheet
The MS sheet lists the observed mass fragments for metabolites. Each row corresponds to a metabolite, with columns for metabolite ID, name, and the mass fragments observed in the experiment.
| Column name | Description |
|---|---|
ID |
Metabolite ID, that must not be duplicated in the model (including model shhet). |
Reaction |
Reaction formula for producing the metabolite to be measured. |
Transition |
Atom mapping information for the reaction. The number of parts must match that of the reaction. Each part describes the mapping from substrate to product. |
Used |
Boolean value indicating whether the mass fragment is used (TRUE) or not (FALSE) in the model fitting. |

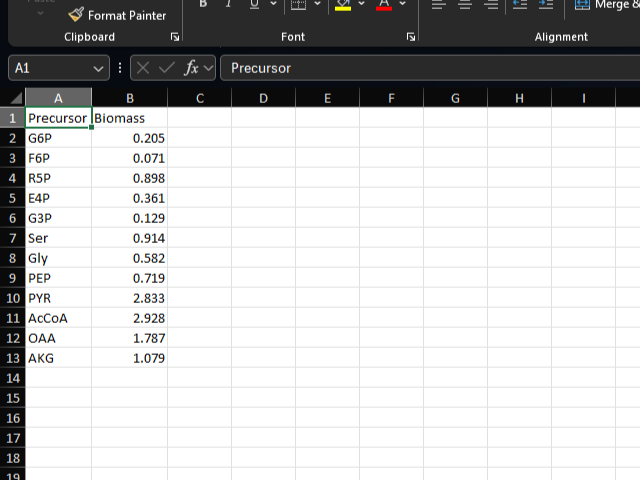
biomass sheet
The biomass sheet defines the biomass coefficients for the biomass equation. Each row represents a metabolite and its coefficient in the biomass equation.
| Column name | Description |
|---|---|
Precursor |
Metabolite name in the biomass equation (must match the metabolite name in the model). |
Biomass |
Coefficient of the metabolite in the biomass equation in mmol gDCW^-1. |
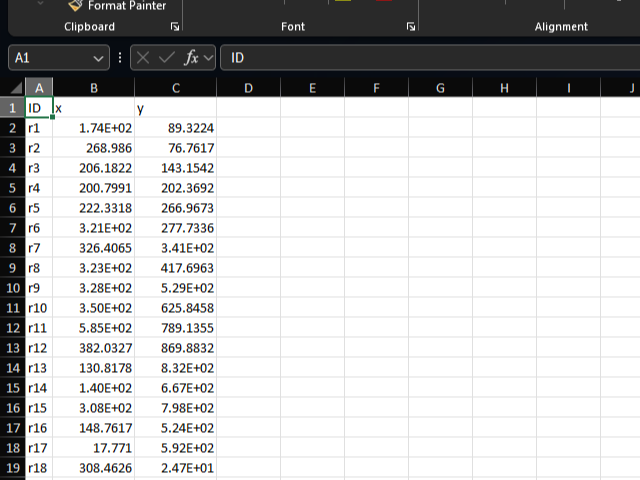
position sheet
The position sheet specifies the positions of reactions in the metabolic map image.
| Column name | Description |
|---|---|
ID |
Reaction ID, that must match the reaction ID in the model sheet. |
x |
X-coordinate of the reaction position in the metabolic map image (in pixels). |
y |
Y-coordinate of the reaction position in the metabolic map image (in pixels). |
label.json file
The label.json file defines the tracer labeling information for each substrate metabolite.
In general, the file is automatically generated when the user inputs tracer information through the GUI.
Each metabolite is represented as a JSON object, with the following structure:
{
"x_1_Glc": {
"name": "[1]Glc",
"num": 6,
"label": [
"#100000",
"#000000"
],
"ratio": [
0.99,
0.01
]
},
"x_12_Glc": {
"name": "[12]Glc",
"num": 6,
"label": [
"#110000",
"#100000",
"#010000",
"#000000"
],
"ratio": [
0.98,
0.01,
0.01,
0
]
},
...
}
| Key | Description |
|---|---|
| Tracer key | Tracer key generated from a tracer name automatically (e.g., x_1_Glc). Edited manually is not recommended. |
name |
Tracer name (e.g., [1]Glc for glucose labeled at carbon 1). name was defined by the user. |
num |
Number of labeled atoms in the metabolite (e.g., 6 for glucose). |
label |
List of labeling patterns represented as binary strings (e.g., #100000 for glucose labeled at carbon 1). |
ratio |
List of fractions corresponding to each labeling pattern. The sum of the ratios must be 1. |